from pathlib import Path
import equinox as eqx
import jax
import jax.numpy as jnp
import matplotlib.pyplot as plt
import numpy as np
import optax
# import scienceplots # noqa: F401
from IPython.display import Audio, display
from jaxtyping import Array, ArrayLike, Float
from tqdm import tqdm
from jaxdiffmodal.excitations import create_pluck_modal
from jaxdiffmodal.ftm import (
StringParameters,
evaluate_string_eigenfunctions,
string_eigenvalues,
)
from jaxdiffmodal.time_integrators import (
solve_sv_ic,
string_tau_with_density,
)
# plt.style.use(["ieee", "no-latex"])
# plt.rcParams["legend.framealpha"] = 1.0
# plt.rcParams["legend.fancybox"] = TrueFitting a synthetic string in time
Utilities
def create_static_filter(
model,
static_params_lambda,
):
is_static_filter = jax.tree_util.tree_map(lambda _: False, model)
selected_params = static_params_lambda(model)
if isinstance(selected_params, tuple):
true_values = tuple(True for _ in selected_params)
else:
# Single parameter case
true_values = True
is_static_filter = eqx.tree_at(
static_params_lambda,
is_static_filter,
true_values,
)
return is_static_filter
def visualize_results(
model,
time: Array,
n_steps_vis: int,
n_steps_train: int,
dt: float,
n_modes: int,
losses: Array | None = None,
):
"""Visualize training results and model predictions."""
print("Generating visualizations...")
time_test = jnp.arange(n_steps_vis) * dt
# Get modal trajectories for visualization
targ_test_traj_modal: Array = gt_model(
n_steps=n_steps_vis,
dt=dt,
n_modes=n_modes,
return_modal=True,
)
pred_test_traj_modal = model(
n_steps=n_steps_vis,
dt=dt,
n_modes=n_modes,
return_modal=True,
)
pred_test_traj_phys: Array = model(
n_steps=n_steps_vis,
dt=dt,
n_modes=n_modes,
return_modal=False,
)
# Target physical position
targ_test_traj_phys: Array = targ_test_traj_position
# Check if losses exist and training is complete
plot_losses = losses is not None and len(losses) > 0
# Create plots - adjust subplot layout based on whether we're plotting losses
if plot_losses:
fig = plt.figure(figsize=(15, 10))
gs_main = fig.add_gridspec(2, 2, hspace=0.3, wspace=0.3)
# Plot loss in bottom right when training is complete
loss_ax = fig.add_subplot(gs_main[1, 1])
loss_ax.semilogy(losses)
loss_ax.set_title("Training Loss")
loss_ax.set_xlabel("Epoch")
loss_ax.set_ylabel("MSE Loss")
loss_ax.grid(True)
else:
fig = plt.figure(figsize=(15, 5))
gs_main = fig.add_gridspec(1, 2, hspace=0.3, wspace=0.3)
# Plot 1: Physical space comparison
physical_ax = fig.add_subplot(gs_main[0, 0])
physical_ax.plot(
time_test[:n_steps_vis],
targ_test_traj_phys[:n_steps_vis],
"b-",
label="Target",
)
physical_ax.plot(
time_test[:n_steps_vis],
pred_test_traj_phys[:n_steps_vis],
"r--",
label="Neural ODE",
)
physical_ax.set_title("Physical Space Displacement")
physical_ax.set_xlabel("Time (s)")
physical_ax.set_ylabel("Displacement (m)")
physical_ax.set_ylim(-0.0025, 0.0025)
physical_ax.grid(True)
physical_ax.axvline(
x=n_steps_train * dt,
color="k",
alpha=1.0,
label="Train/Test Split",
)
physical_ax.legend(loc="upper right")
# Plot 2: Modal amplitudes comparison - use the right half of top row
if plot_losses:
# Create subplot within the top-right area, avoiding the loss plot
modal_gs = gs_main[0, 1].subgridspec(3, 1, hspace=0.4)
else:
# Use the right half for modal plots
modal_gs = gs_main[0, 1].subgridspec(3, 1, hspace=0.4)
for mode_idx in range(min(3, n_modes)):
modal_ax = fig.add_subplot(modal_gs[mode_idx, 0])
modal_ax.plot(
time_test,
targ_test_traj_modal[:n_steps_vis, mode_idx],
label="Target",
alpha=0.8,
linewidth=1.5,
)
modal_ax.plot(
time_test,
pred_test_traj_modal[:n_steps_vis, mode_idx],
"--",
label="Prediction",
alpha=0.8,
linewidth=1.5,
)
modal_ax.set_title(f"Mode {mode_idx + 1}", fontsize=10)
if mode_idx == 2: # Only bottom plot gets x-label
modal_ax.set_xlabel("Time (s)", fontsize=9)
modal_ax.set_ylabel("Amplitude", fontsize=9)
modal_ax.set_ylim(-0.0083, 0.0083)
modal_ax.tick_params(labelsize=8)
if mode_idx == 0: # Only top plot gets legend
modal_ax.legend(fontsize=8)
modal_ax.grid(True, alpha=0.3)
modal_ax.axvline(
x=n_steps_train * dt,
color="k",
alpha=0.7,
linestyle=":",
)
plt.show()Step 1: Generate Synthetic String Data
First, we’ll create synthetic string data using jaxdiffmodal’s physical model. We’ll generate both linear and nonlinear dynamics to have target data for training.
n_modes: int = 15
sample_rate: int = 16000
dt: float = 1.0 / sample_rate
n_steps_train: int = 1000
n_steps_test: int = 16000
n_steps_vis = 2000string_params = StringParameters()
indices = jnp.arange(n_modes) + 1
lambda_mu = string_eigenvalues(
n_modes,
string_params.length,
)
exc = create_pluck_modal(
lambdas=lambda_mu,
string_length=string_params.length,
initial_deflection=0.03,
)
weights = evaluate_string_eigenfunctions(
indices=indices,
position=jnp.array(0.6),
params=string_params,
)
u0 = jnp.array(exc)
v0 = jnp.zeros_like(u0)
time = jnp.arange(n_steps_train) * dt
class StringModel(eqx.Module):
length: ArrayLike
d3_with_density: ArrayLike
log_Ts0_with_density: ArrayLike
bending_stiffness_with_density: ArrayLike
tau_with_density: ArrayLike
v0: Array
u0: Array
weights: Array # Modal weights for single position output
mlp: eqx.Module | None = None
def __call__(
self,
n_steps: int,
dt: float,
n_modes: int = 10,
return_modal: bool = False,
) -> Float[Array, " n_steps"] | Float[Array, "n_steps n_modes"]:
# Unpack parameters
length: ArrayLike = self.length
d3_with_density: ArrayLike = self.d3_with_density
# Convert from log-space
Ts0_with_density: ArrayLike = jnp.exp(self.log_Ts0_with_density)
bending_stiffness_with_density: ArrayLike = self.bending_stiffness_with_density
tau_with_density: ArrayLike = self.tau_with_density
u0: Array = self.u0
v0: Array = self.v0
# get the analytical eigenvalues
lambda_mu: Array = string_eigenvalues(
n_modes,
length,
)
# get the damping and stiffness terms
omega_mu_squared: Array = (
bending_stiffness_with_density * lambda_mu**2 + Ts0_with_density * lambda_mu
)
gamma2_mu: Array = d3_with_density * lambda_mu
# calculate the factor for the nonlinear term
string_norm: float = string_params.length / 2
string_tau: Array = tau_with_density * lambda_mu / string_norm
def nl_fn(q: ArrayLike) -> Array:
return lambda_mu * q * (string_tau @ q**2)
def nl_fn_nn(q: ArrayLike) -> Array:
return lambda_mu * self.mlp(q)
_, traj = solve_sv_ic(
gamma2_mu=gamma2_mu,
omega_mu_squared=omega_mu_squared,
u0=u0,
v0=v0,
dt=dt,
n_steps=n_steps,
nl_fn=nl_fn_nn if self.mlp is not None else nl_fn,
)
if return_modal:
return traj
else:
# Apply weights to get single position output
return traj @ self.weightsstring_tau: float = string_tau_with_density(string_params)
gt_model = StringModel(
length=string_params.length,
log_Ts0_with_density=jnp.log(string_params.Ts0 / string_params.density),
d3_with_density=(string_params.d3 / string_params.density),
bending_stiffness_with_density=(
string_params.bending_stiffness / string_params.density
),
tau_with_density=string_tau_with_density(string_params),
u0=u0,
v0=v0,
weights=weights,
mlp=None,
)
# Get weighted trajectory at single position for training target
targ_test_traj_position: Array = gt_model(
n_steps=n_steps_test,
dt=dt,
n_modes=n_modes,
return_modal=False,
)
# slice a section for training
targ_train_traj_position: Array = targ_test_traj_position[:n_steps_train]# Initialize model with random weights for optimization
key = jax.random.PRNGKey(12345)
(
key_len,
key_Ts0,
key_d3,
) = jax.random.split(key, 3)
model = StringModel(
length=jax.random.uniform(
shape=(1,),
minval=0.6,
maxval=0.8,
key=key_len,
),
log_Ts0_with_density=jax.random.uniform(
shape=(1,),
minval=jnp.log(10_000),
maxval=jnp.log(80_000),
key=key_Ts0,
),
d3_with_density=jax.random.uniform(
shape=(1,),
minval=5.0,
maxval=7.0,
key=key_d3,
),
bending_stiffness_with_density=(
string_params.bending_stiffness / string_params.density
),
tau_with_density=string_tau_with_density(string_params),
u0=u0,
v0=v0,
weights=weights,
mlp=None,
)
# Create the static filter using the wrapper function
is_static_filter = create_static_filter(
model=model,
static_params_lambda=lambda m: (
m.v0,
m.u0,
m.weights,
m.bending_stiffness_with_density,
),
)
# Now, partition the model using our custom filter
static_model, diff_model = eqx.partition(
model,
is_static_filter,
)
pred_init_traj_position = model(
n_steps=n_steps_test,
dt=dt,
n_modes=n_modes,
return_modal=False,
)
display(Audio(targ_test_traj_position, rate=sample_rate))
display(Audio(pred_init_traj_position, rate=sample_rate))Define the training loop and loss function.
def save_animation_frame(
model,
time: Array,
weights: Array,
frame_idx: int,
gt_model,
output_dir: str = "tmp_node",
):
Path(output_dir).mkdir(exist_ok=True, parents=True)
time_test = jnp.arange(n_steps_test) * dt
pred_test_traj_modal = model(
n_steps=n_steps_test,
dt=dt,
n_modes=n_modes,
return_modal=True,
)
pred_test_traj_phys: Array = model(
n_steps=n_steps_test,
dt=dt,
n_modes=n_modes,
return_modal=False,
)
# Target physical position
targ_test_traj_phys: Array = targ_test_traj_position
# Create figure with centered plot and table underneath
fig = plt.figure(figsize=(12, 6))
gs = fig.add_gridspec(2, 1, height_ratios=[4, 1], hspace=0.4)
# Plot 1: Physical space comparison (centered)
physical_ax = fig.add_subplot(gs[0, 0])
physical_ax.plot(
time_test[: n_steps_vis * 2],
targ_test_traj_phys[: n_steps_vis * 2],
"b-",
label="Target",
)
physical_ax.plot(
time_test[: n_steps_vis * 2],
pred_test_traj_phys[: n_steps_vis * 2],
"r--",
label="Optim",
)
physical_ax.set_title("Physical Space Displacement")
physical_ax.set_xlabel("Time (s)")
physical_ax.set_ylabel("Displacement (m)")
physical_ax.set_ylim(-0.0025, 0.0025)
physical_ax.legend(loc="upper right")
physical_ax.grid(True)
physical_ax.axvline(
x=n_steps_train * dt,
color="k",
linestyle=":",
alpha=0.7,
label="Train/Test Split",
)
# Add parameter table
table_ax = fig.add_subplot(gs[1, 0])
table_ax.axis("off")
# Create table data - handle both JAX arrays and floats
def format_param(param):
return param.item() if hasattr(param, "item") else param
table_data = [
["Parameter", "Current", "Ground Truth"],
[
"Length",
f"{format_param(model.length):.4f}",
f"{format_param(gt_model.length):.4f}",
],
[
r"$\hat{d}_3$",
f"{format_param(model.d3_with_density):.6f}",
f"{format_param(gt_model.d3_with_density):.6f}",
],
[
r"$\hat{T}_0$",
# Show actual value, not log
f"{format_param(jnp.exp(model.log_Ts0_with_density)):.1f}",
f"{format_param(jnp.exp(gt_model.log_Ts0_with_density)):.1f}",
],
]
table = table_ax.table(
cellText=table_data,
cellLoc="center",
loc="center",
colWidths=[0.25, 0.25, 0.25],
)
table.auto_set_font_size(False)
table.set_fontsize(12)
table.scale(1, 2)
# Style the header row
for i in range(len(table_data[0])):
table[(0, i)].set_facecolor("#40466e")
table[(0, i)].set_text_props(weight="bold", color="white")
plt.tight_layout()
plt.savefig(f"{output_dir}/frame_{frame_idx:05d}.png", dpi=150, bbox_inches="tight")
plt.close()def train_neural_ode(
model,
save_frames=False,
frame_interval=50,
):
print("Training...")
# normalise target trajectory for better training stability
# Now using single position target instead of modal trajectories
scale: float = jnp.max(jnp.abs(targ_train_traj_position)).item()
targ_train_traj_position_scaled = targ_train_traj_position / scale
@eqx.filter_jit
def training_step(
model,
optimizer,
opt_state,
targ_train_traj_position_scaled,
):
@eqx.filter_value_and_grad
def loss_fn(
diff_model,
static_model,
targ_train_traj_position_scaled,
):
model: eqx.Module = eqx.combine(diff_model, static_model)
pred_train_traj_position: Array = model(
n_steps=n_steps_train,
dt=dt,
n_modes=n_modes,
return_modal=False,
)
pred_train_traj_position: Array = (
pred_train_traj_position / scale
) # normalise predictions
# MSE loss
mse_loss = jnp.mean(
(pred_train_traj_position - targ_train_traj_position_scaled) ** 2
)
total_loss = mse_loss
return total_loss
static_model, diff_model = eqx.partition(
model,
is_static_filter,
)
loss_value, grads = loss_fn(
diff_model,
static_model,
targ_train_traj_position_scaled,
)
updates, opt_state = optimizer.update(grads, opt_state)
model = eqx.apply_updates(model, updates)
return model, opt_state, loss_value
# Training setup
epochs = 10000
learning_rate = 1e-4
schedule = optax.cosine_onecycle_schedule(
transition_steps=epochs,
peak_value=learning_rate,
)
optimizer = optax.chain(
optax.clip_by_global_norm(1.0),
optax.adabelief(schedule),
)
opt_state = optimizer.init(
eqx.filter(model, eqx.is_array),
)
losses = []
bar = tqdm(range(epochs))
for epoch in bar:
model, opt_state, loss_value = training_step(
model,
optimizer,
opt_state,
targ_train_traj_position_scaled,
)
losses.append(loss_value)
# Early stopping if NaN detected or loss explodes
if jnp.isnan(loss_value) or loss_value > 1e8:
print(
f"\nWarning: Training stopped early at epoch {epoch + 1} due to instability"
)
print(f"Loss value: {loss_value}")
break
bar.set_description(f"Epoch {epoch + 1}/{epochs} | Loss: {loss_value:.6f}")
# Save animation frame periodically
if save_frames and epoch % frame_interval == 0:
save_animation_frame(
model=model,
time=time,
weights=weights,
frame_idx=epoch // frame_interval,
gt_model=gt_model,
)
return model, losses# First visualisation of the initial model
visualize_results(
model=model,
time=time,
n_steps_vis=n_steps_vis,
n_steps_train=n_steps_train,
dt=dt,
n_modes=n_modes
)Generating visualizations...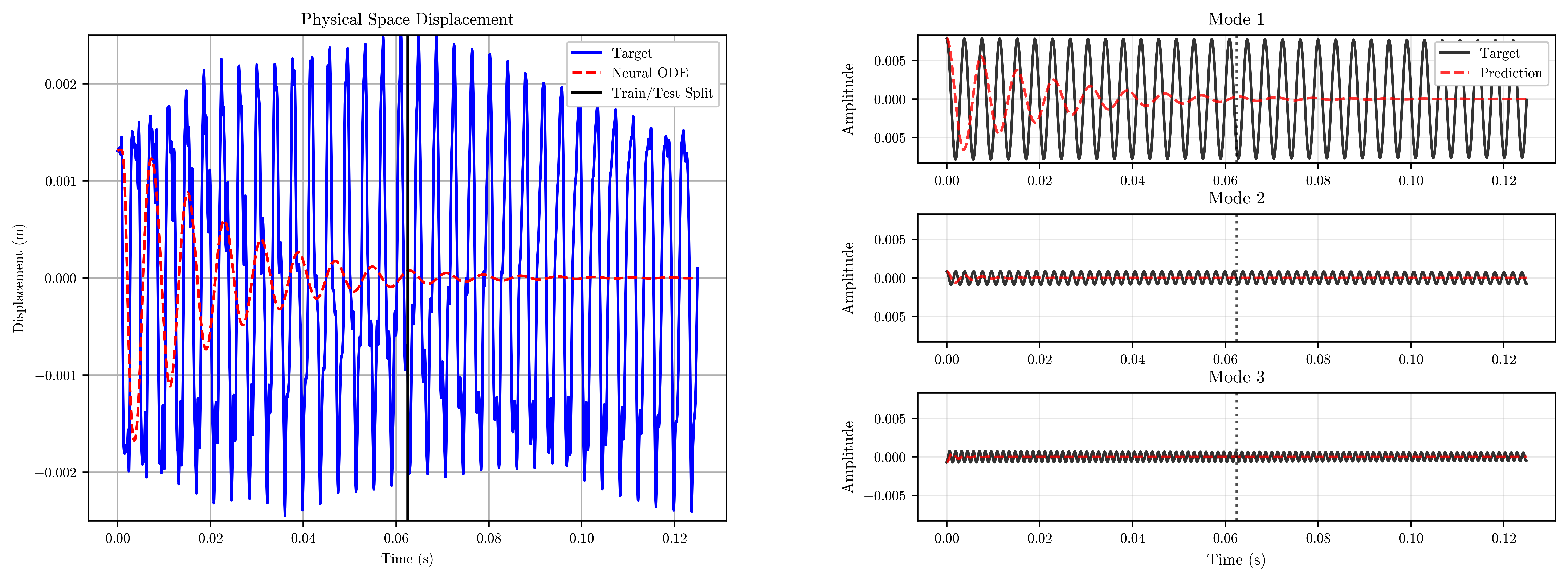
print("Starting training...")
trained_model, training_losses = train_neural_ode(
model,
save_frames=False,
frame_interval=100,
)
print(f"Training completed! Final loss: {training_losses[-1]:.6f}")Starting training...
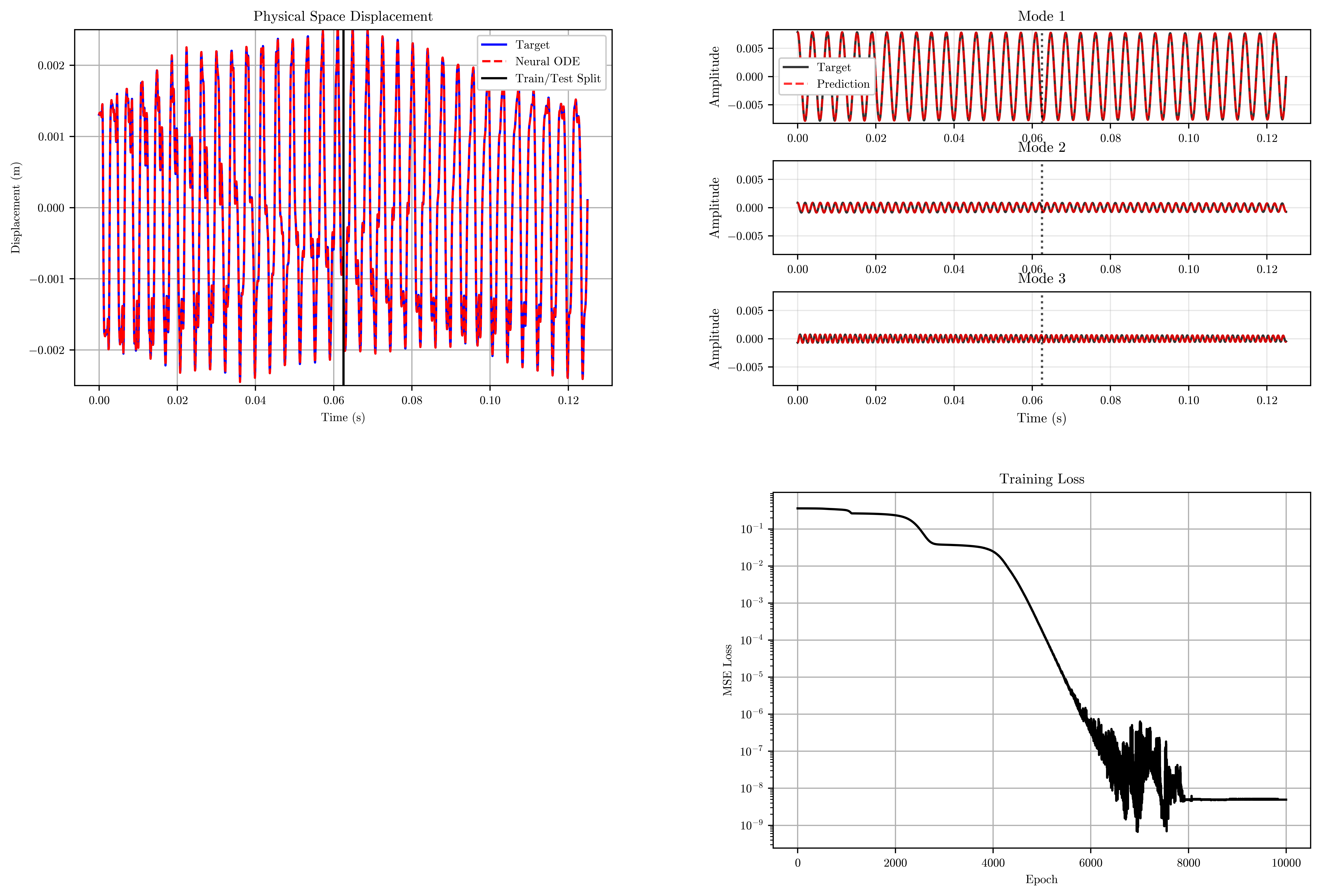
Training...Epoch 10000/10000 | Loss: 0.000000: 100%|██████████| 10000/10000 [00:24<00:00, 402.16it/s]Training completed! Final loss: 0.000000visualize_results(
model=trained_model,
time=time,
losses=training_losses,
n_steps_vis=n_steps_vis,
n_steps_train=n_steps_train,
dt=dt,
n_modes=n_modes
)Generating visualizations...